Fragment Based Drug Design
Thinking of fragment derivatives is easy: the challenge is to automate the process of generating quality derivatives and evaluating their binding potential. Read our white paper De Novo and Combichem Growth using THINK for Fragment Based Drug Design.
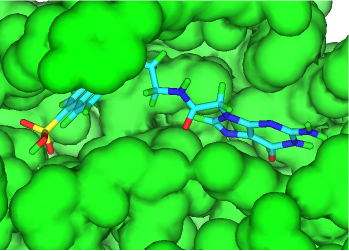
The de novo derivative generation and Combichem enumeration functionality of THINK have been enhanced to provide in situ growth of docked fragments to more strongly binding molecules which interact with targeted residues whilst retaining their drug-like characteristics. Critical algorithms have been improved and the number of transformations which can be applied to molecules has been increased. The software integration within THINK allows these poses to be refined with protein side-chain motion and ranked using a scoring function.
The FBDD capabilities can be used in stand-alone mode or integrated into the KNIME environment. More information about how to use these capabilities can be found in the release notes.
Can you help with our P450 project?
Our marathon P450 inhibition project which started over 10 years ago has reached a significant milestone and later this year you will be able to select molecules for purchase or synthesis with 90% confidence of avoiding P450 inhibition. One more way with THINK that you can avoid optimising leads which would subsequently fail in clinical trials. Can you spare 2 minutes to complete our online survey?
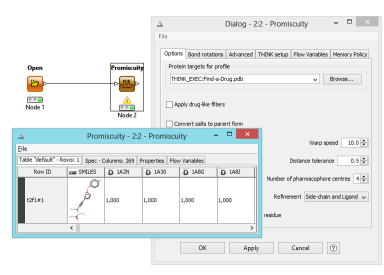
THINK 2.0 released
Early this summer THINK 2.0 was released. This release includes a significant rewrite of a number of key algorithms used by the 3D pharmacophore technology in order to take better advantage of CPU instruction sets to improve performance especially when refining protein side-chain orientations. These changes were important for the new FBDD capabilities as well as a new Promiscuity KNIME node which allows molecules to be effortlessly profiled across hundreds of protein targets to evaluate their selectivity and propensity to be promiscuous.